Our client was developing novel compounds targeting a metabolic gatekeeper protein. The existing screening cascade was based on a complex in vitro evaluation assay utilising isolated mitochondria, which was low-throughput, labour-intensive, technically demanding and susceptible to variability from biological sample preparation.
Mitochondrial biology experts at BioAscent developed a customised bioenergetic workflow using the Seahorse XF Pro platform. The assay was optimised to mitigate the risk of compound penetrance issues and plasma membrane-level off-target effects while producing robust data. Additional mechanism of action studies confirmed on-target specificity.
Our optimised assay cascade reliably generated precise efficacy profiles and distinguished genuine on-target effects from off-target toxicity. It successfully eliminated a critical testing bottleneck and gave the client’s scientific team the confidence to rapidly interpret emerging SAR and accelerate their design-make-test-analyse cycles.
Cellular bioenergetics relies on the tightly regulated transport of metabolites across the mitochondrial membrane. The target of this project is a critical metabolic gatekeeper protein that controls the influx of a primary carbon substrate into the mitochondrial matrix, dictating the rate of the tricarboxylic acid (TCA) cycle and fundamentally sustaining oxidative phosphorylation (OXPHOS).
The dysregulation of this gatekeeper protein is a driving factor in several complex pathologies, including oncological diseases and systemic metabolic disorders. By blocking this crucial metabolic entry point, it is possible to selectively disrupt the altered energy metabolism of diseased cells or force a beneficial metabolic reprogramming, making it an attractive target for novel drug discovery programmes.
Apollo Therapeutics, a UK-US biopharmaceutical company developing innovative new medicines, required a robust, high-throughput functional assay to rapidly and reliably screen novel compounds targeting this metabolic gatekeeper.
Investigating specific mitochondrial-related targets presents unique challenges in early-stage drug discovery. Designing and interpreting mitochondrial functional assays is inherently complex due to the organelle's double-membrane architecture and its highly adaptable, interconnected metabolic networks. In intact cellular models, compensatory metabolic routing and variable compound penetrance across the plasma membrane can easily mask the direct pharmacological effects on an inner-membrane target. Conversely, isolating mitochondria to bypass these cellular barriers strips away vital physiological context and introduces significant handling artefacts.
Apollo’s existing screening cascade was based on traditional gold-standard methodology which relied on a complex in vitro evaluation assay utilising isolated mitochondria. While the assay provided a direct mechanistic readout, it was:
Labour-intensive
Technically demanding
Inherently low-throughput
Susceptible to variability from biological sample preparation
BioAscent’s mitochondrial biology experts sought to overcome this limitation by designing a sensitive, highly reproducible, and robust functional bioassay that could accurately capture the functional inhibition of this specific gatekeeper in a format scalable for high-throughput screening, while maintaining the mechanistic confidence provided by isolated organelle systems.
A customised bioenergetic workflow was developed using the Seahorse XF Pro platform. Assay optimisation focused on isolating the precise metabolic contribution of the target gatekeeper to mitochondrial function through four key strategic steps:
1 - Assay streamlining and variability reduction
We developed a refined version of the canonical Seahorse Mito Stress Test. By redesigning the port injection strategy to capture only the essential metrics reflecting the target protein's respiration readout, unnecessary complexity was removed from the assay (Figure 1). This streamlined approach significantly reduced data variability and provided the assay robustness required for high-throughput screening.
2 - Metabolic isolation via medium optimisation
To ensure the respiratory signal was completely dependent on the target, we carefully optimised the assay medium. The composition was tailored to restrict metabolic fuel exclusively to the substrates associated with this specific membrane gatekeeper, forcing the bioenergetic output to act as a direct proxy for its function.
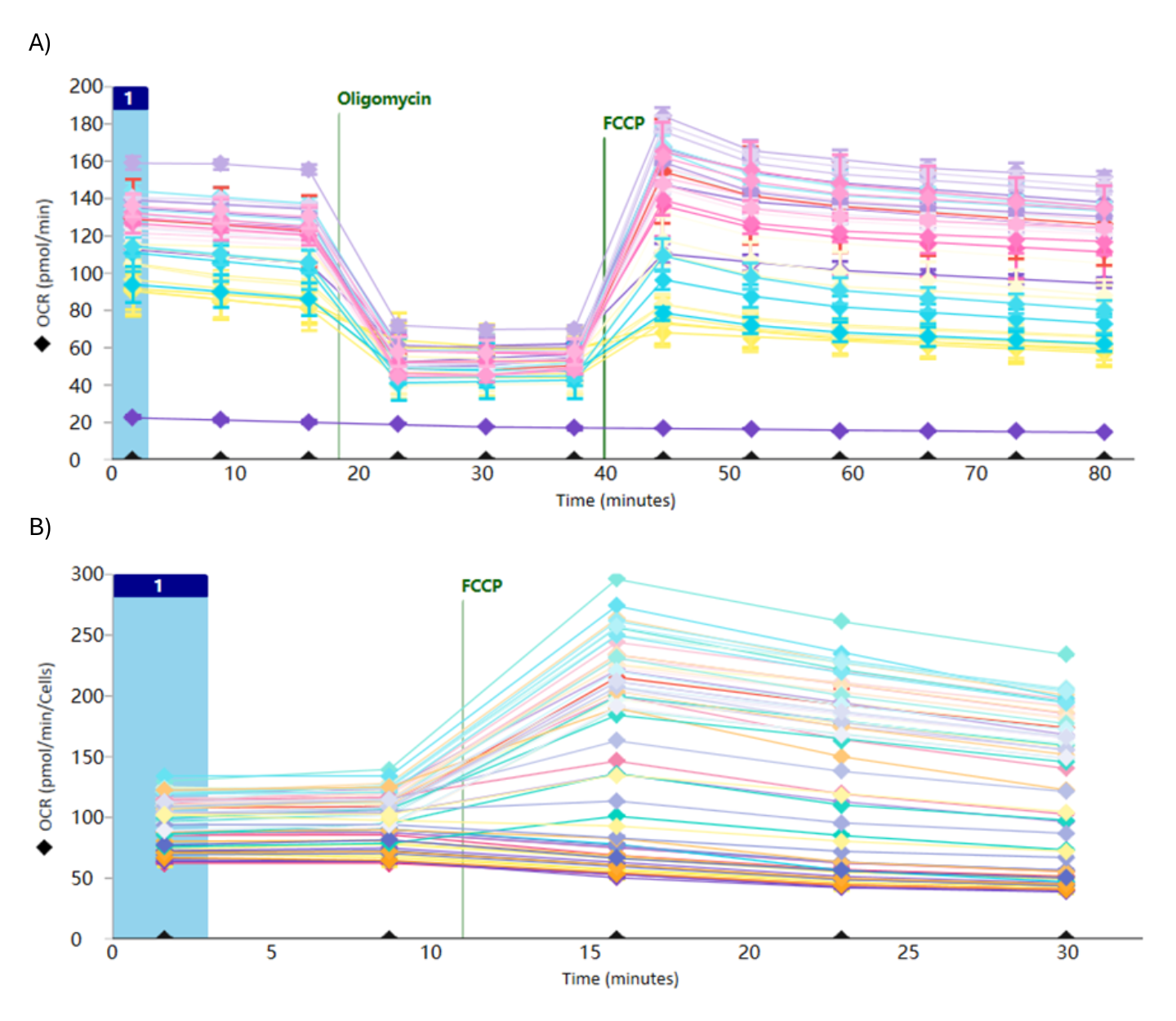
Figure 1. During initial assay development, compounds were pretreated with the oxidative substrate (alpha) and evaluated using a 10-point dose–response curve (DRC). Two experimental strategies were employed: A) sequential port-injection of oligomycin followed by FCCP, and B) FCCP injection alone. These approaches were designed to examine potential inhibition of oxygen consumption rate (OCR), elucidate the extent to which the compounds perturb specific pathways of OXPHOS, and reduce the assay variability. FCCP = Carbonyl cyanide-p-trifluoromethoxyphenylhydrazone), a potent mitochondrial oxidative phosphorylation uncoupler.
3 - Direct mitochondrial targeting via permeabilisation
The target protein is localised to the mitochondrial membrane; therefore, intact cellular models pose a risk of compound penetrance issues or plasma membrane-level off-target effects. To circumvent this, we utilised Plasma Membrane Permeabilizer (PMP). By meticulously optimising the formula of the permeabilisation media (MAS buffer), we successfully exposed the mitochondria directly to the test compounds in situ. This bridged the gap between high-throughput cellular assays and the traditional isolated-mitochondria gold standard (Figure 2; Figure 3).
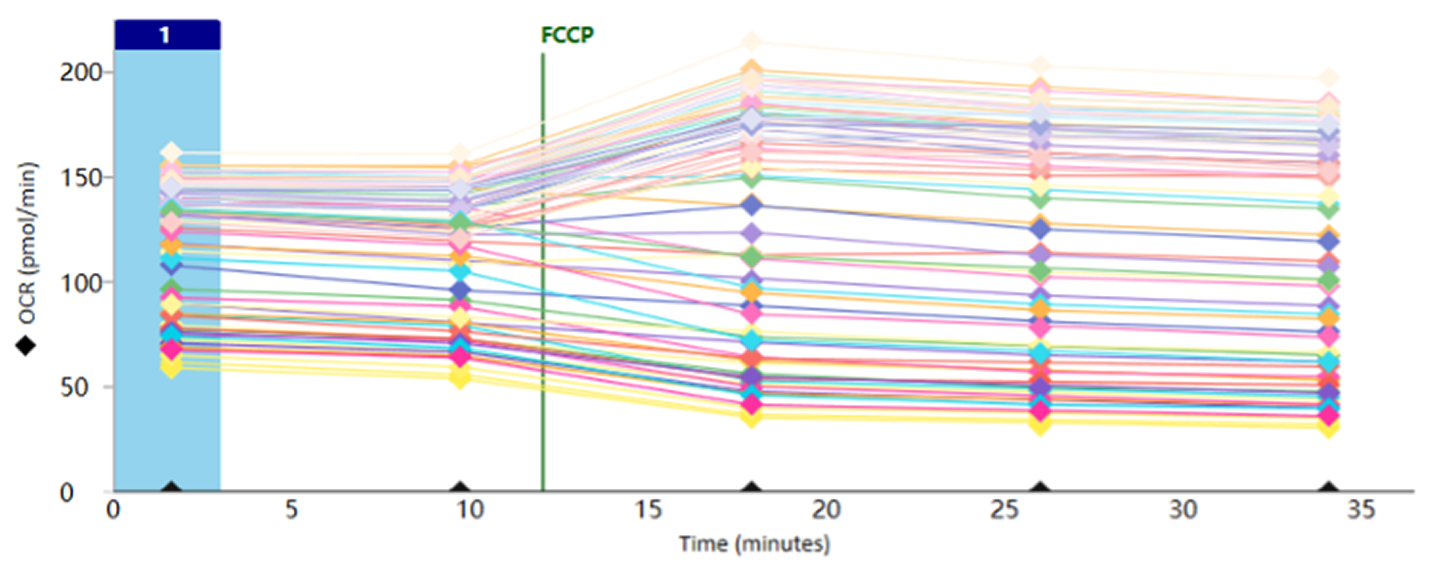
Figure 2. The optimised assay conditions yielded robust OCR profiles in permeabilised HepG2 cells (10-pt DRC).
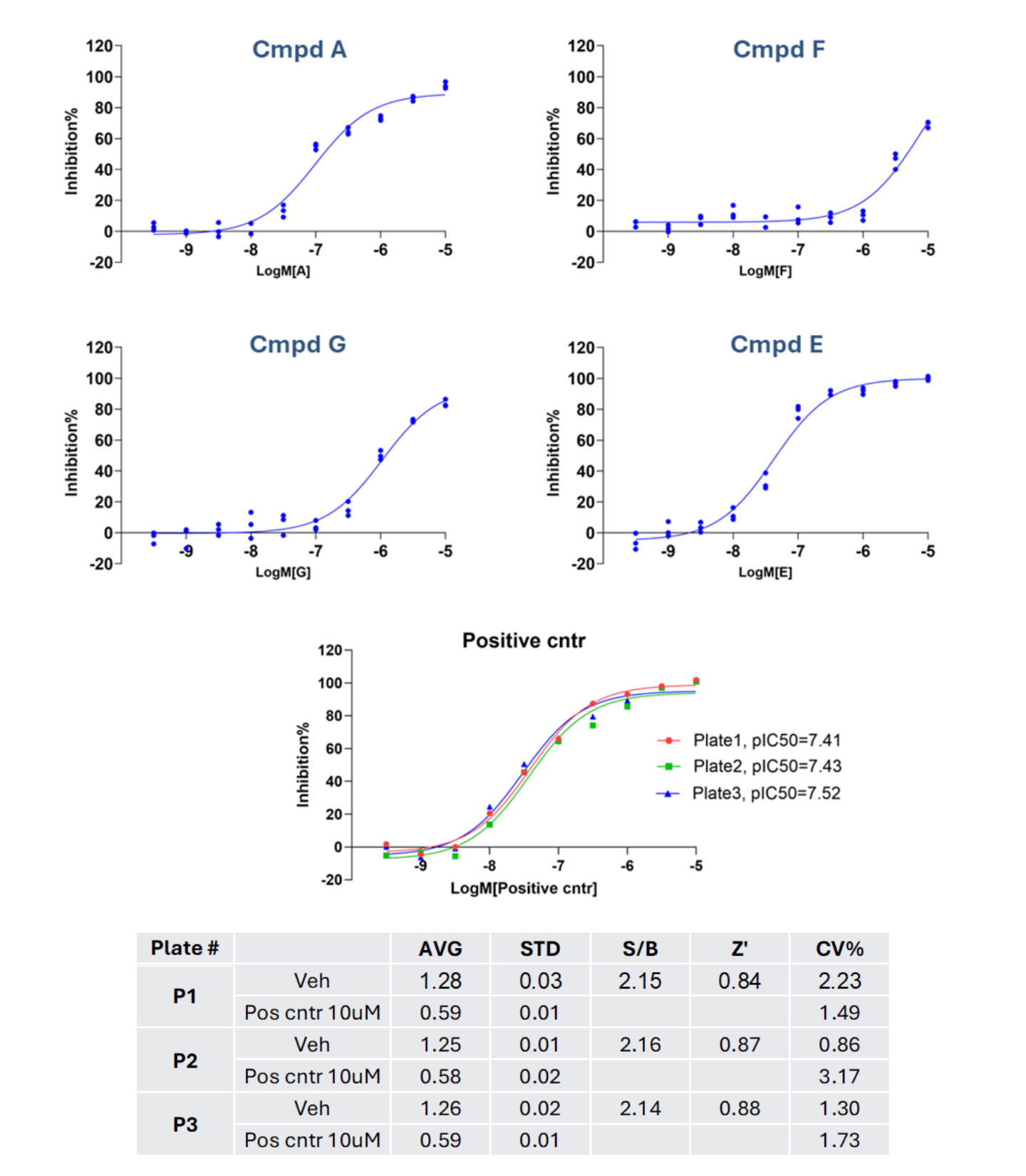
Figure 3. Screening results demonstrated that the tested compounds displayed distinct potencies relative to the intra-plate positive control (a specific inhibitor of ‘X’ signalling), and the assay exhibited a high degree of robustness.
4 - Orthogonal validation of specificity
To provide definitive proof of mechanism, we engineered a customised Seahorse orthogonal counter-screen. Permeabilised HepG2 cells were supplied with a unique alternative substrate designed to metabolically bypass the target gatekeeper. By demonstrating that compound-induced respiratory inhibition was attenuated or fully abolished under these bypass conditions, we could confidently confirm on-target specificity for each compound (Figure 4).
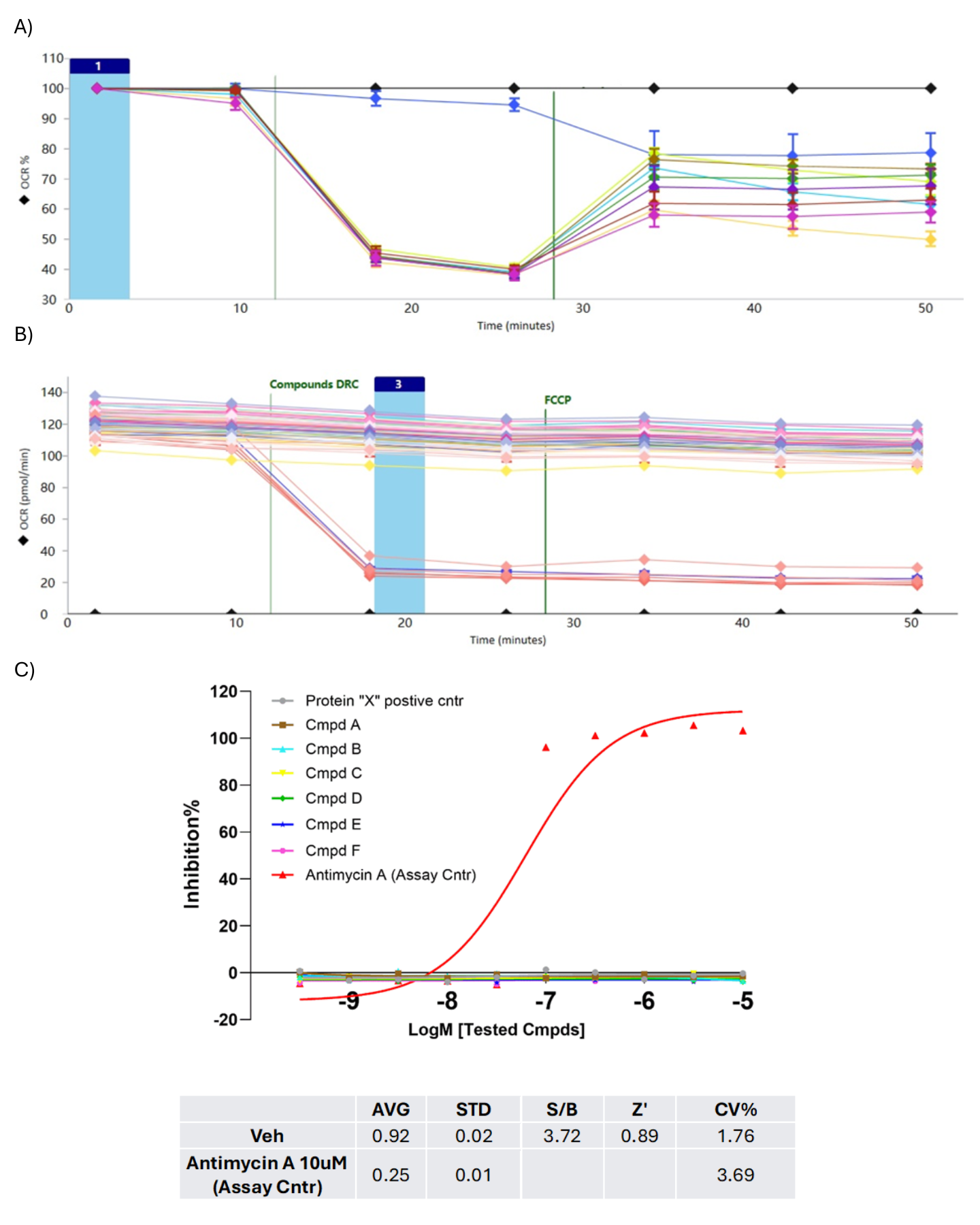
Figure 4. For orthogonal assay development, A) the optimised concentration of an alternative oxidative substrate (beta), acting through the target gatekeeper protein ‘X’ signalling pathway, was titrated to enable further characterisation. B) Hit compounds were evaluated using an optimised orthogonal assay to assess their specificity. Following port-injection of the hits and the assay control (antimycin A), FCCP was injected to induce mitochondrial uncoupling. C) The resulting data indicated that the compounds exhibited high specificity toward the target gatekeeper protein ‘X’ in its signalling. The assay demonstrated strong robustness and reproducibility.
BioAscent scientists successfully delivered a reliable, high-throughput alternative to traditional, labour-intensive isolated mitochondria assays.
By bridging the gap between intact cellular models and isolated organelle systems, the optimised assay reliably generated precise efficacy profiles while effectively distinguishing genuine on-target compounds from those exhibiting off-target toxicity.
Our optimised screening pipeline eliminated a critical testing bottleneck and gave the scientific team at Apollo confidence to rapidly interpret emerging SAR and accelerate their design-make-test-analyse cycles.
Discover more about BioAscent’s mitochondrial biology expertise here.
The work in this case study was performed by BioAscent Senior Scientist Yuxin Wu.
Yuxin leads mitochondrial research projects at BioAscent and has extensive experience in the use of Seahorse bioenergetic assays to address complex scientific questions.





