High-throughput screening (HTS) remains a central technique for early drug discovery, but compounds that trigger assay responses through non-specific, non‑mechanistic, or artefactual modes of action frequently contaminate hit lists. Left unchecked, these misleading actives can drain resources, misdirect teams, and significantly delay programme progression.
At BioAscent we routinely employ our proprietary PAINS (Pan Assay INterference compoundS) library during assay development to systematically optimise assay buffer conditions, minimise false positives and increase our probability of identifying genuine hits. This curated collection contains ~150 compounds representing major interference mechanisms including aggregators, chelators, coloured/fluorescent compounds, redox cyclers, luciferase inhibitors, reactive molecules, and synthesis contaminants (Figure 1).
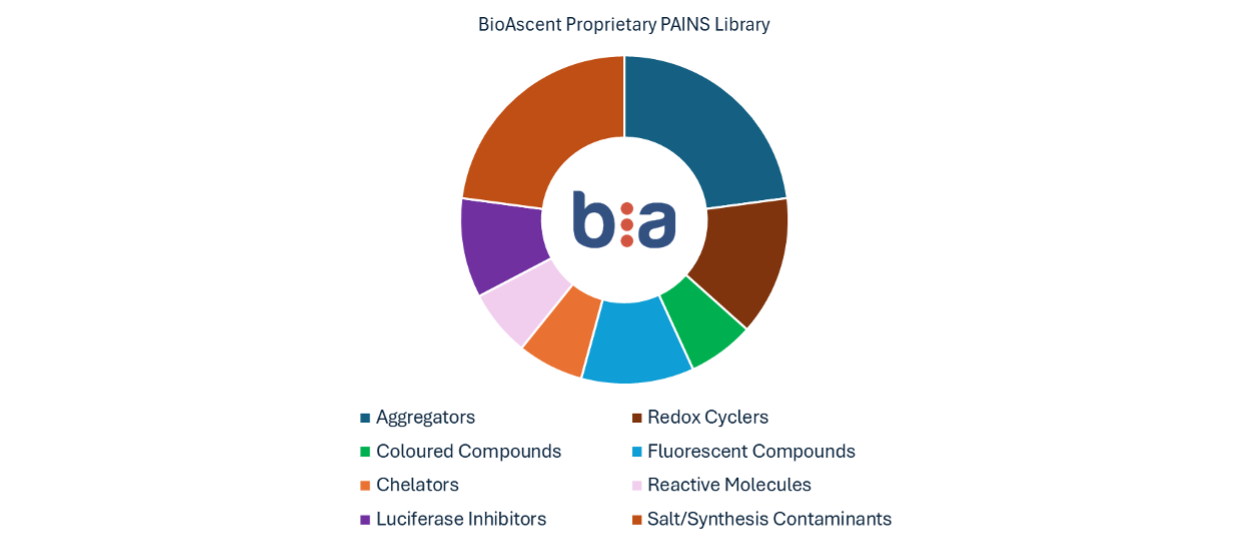
Figure 1. Composition of BioAscent’s PAINS library.
BioAscent scientists were working to develop a Fluorescence Intensity (FI)-based assay for an HTS against a metalloenzyme target. PAINS library compounds were tested at 10 µM (n = 4) during assay development using the original buffer (20 mM HEPES pH 7.5, 100 mM KCl, 1.5 mM MgCl2, 0.02% NP-40, 3% glycerol, 0.5 mM DTT, 0.1 mg/mL BSA). Many compounds showed >20% inhibition suggesting the target was highly susceptible to interference by PAINS compounds under these conditions (Figure 2).
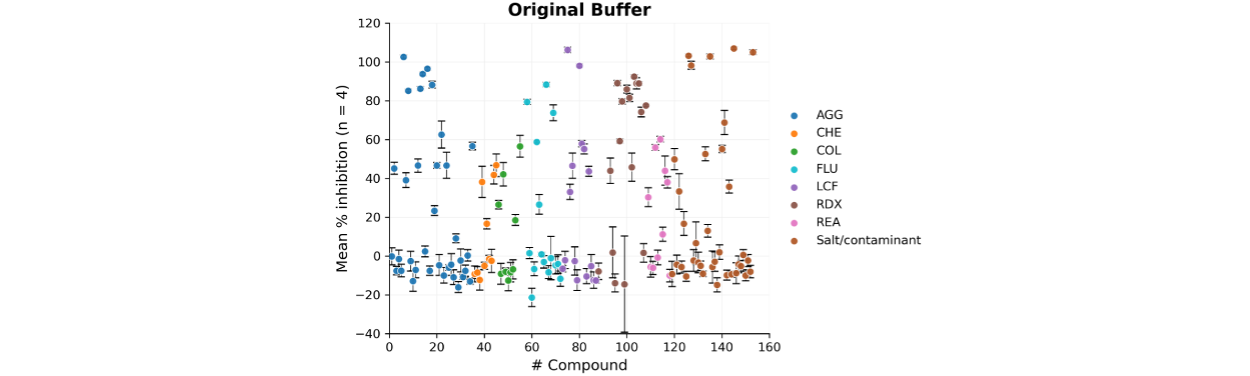
Figure 2. Scatter plot showing the mean normalised % inhibition (n = 4) for each compound in the original assay buffer, calculated using the maximum (Reference compound) and minimum (DMSO) controls. Error bars represent the standard error of the mean (SEM).
Strategic Buffer Optimisation to Reduce Artefacts
Early profiling of the target with a representative panel of PAINS compounds rapidly revealed significant vulnerabilities in the assay. Drawing on knowledge of the target’s biochemistry, our extensive experience characterising PAINS sensitivity across hundreds of proteins, and the specific PAINS classes to which this target appeared most susceptible, we then refined the assay to minimise or eliminate sensitivity to the predicted interference mechanisms.
To achieve this, we systematically evaluated the influence of key buffer components, including reducing agents, detergents, and metal chelators, on the behaviour of PAINS-like actives. Each buffer variable was adjusted individually, and all compounds were tested in duplicate or quadruplicate to ensure robustness and reproducibility of the resulting data.
Effect of DTT
DTT is a reducing agent that helps protect the enzyme from oxidative damage, which can alter its structure and impair activity. Certain PAINS compounds can cause oxidation reactions that damage the enzyme function, which can be mitigated by strong reducing agents such as DTT or TCEP. An important caveat is that some compounds can themselves react with strong reducing agents to produce radicals such as peroxide or superoxide to paradoxically enhance unwanted enzyme oxidation.
We found that increasing DTT from 0.5 mM to 5 mM provided some modest reduction in PAINS related interference across a range of compound classifications, including redox-active compounds (Figure 3).
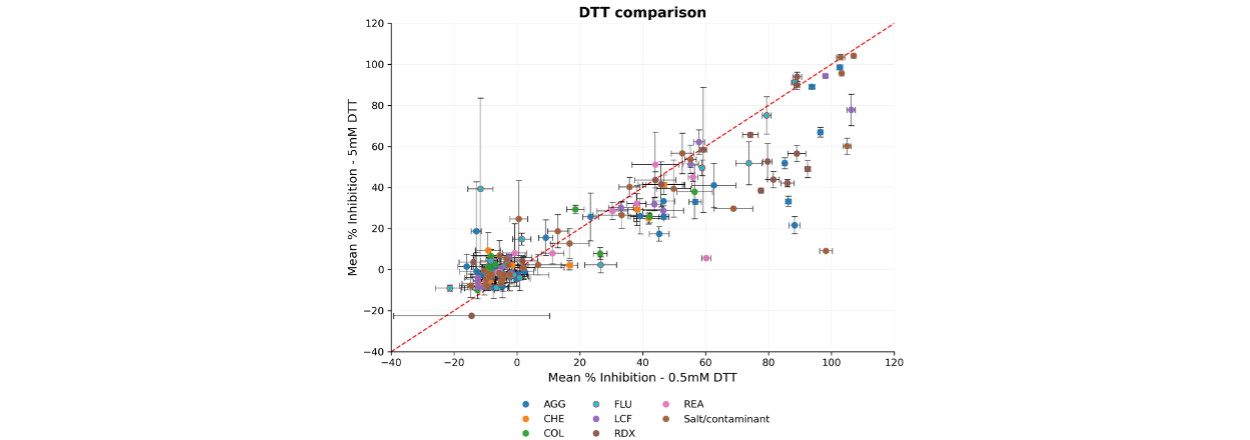
Figure 3. Comparison of DTT levels in assay buffer composition. Correlation plot showing the mean normalised % inhibition (n = 2/4) for each compound, calculated using the maximum (Reference compound) and minimum (DMSO) controls. The red dashed line represents the line of unity (y = x). Error bars represent the standard error of the mean (SEM).
Introduction of EGTA
EGTA is a divalent metal chelator that effectively sequesters contaminating cations, preventing them from interacting with assay components, including the enzyme. In many screening libraries residual transition metals such as Zn²+ or Pd²+, commonly introduced during catalytic synthetic steps, can persist at low levels and contribute to assay interference. Because the target enzyme requires magnesium to maintain activity, EGTA was selected over EDTA due to its substantially lower affinity for Mg²⁺. This allows EGTA to chelate unwanted contaminant metals without stripping the essential magnesium cofactor from the enzyme.
Incorporating 5 mM EGTA into the assay buffer significantly reduced spurious inhibition, with activity suppression falling below 20% across most PAINS classes (Figure 4).
It is also important to recognise that some PAINS chemotypes can act as chelators themselves. These structures may bind essential metal ions or interact with residual catalytic metals, altering cofactor availability and generating misleading signals, either by artificially enhancing apparent activity or producing non-specific inhibition. Such metal‑dependent artefacts illustrate the need for careful buffer optimisation and mechanistic follow‑up when interpreting early screening hits from chemically reactive or interference‑prone scaffolds.
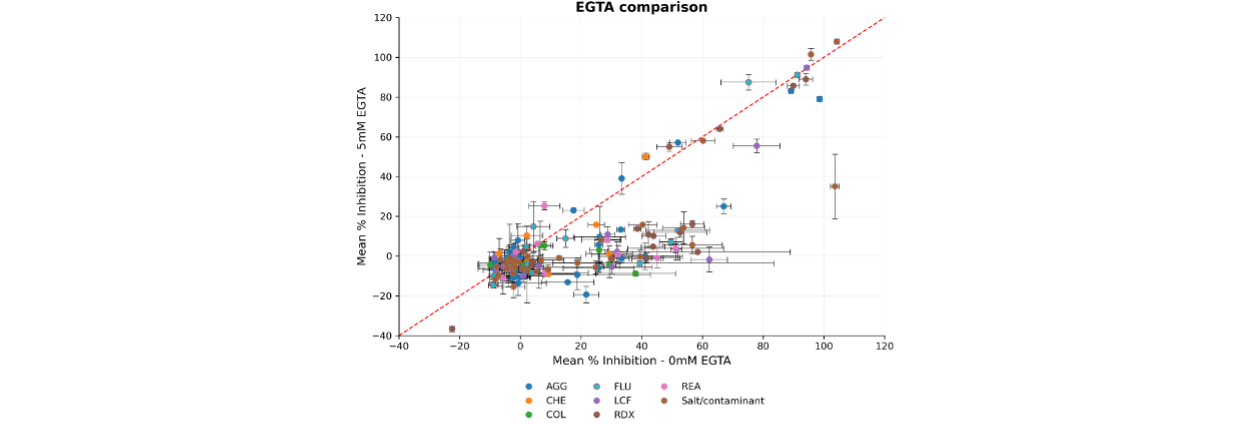
Figure 4. Comparison of assay performance with and without the inclusion of 5 mM EGTA in the buffer. Correlation plot showing the mean normalised % inhibition (n = 4) for each compound, calculated using the maximum (Reference compound) and minimum (DMSO) controls. The red dashed line represents the line of unity (y = x). Error bars represent the standard error of the mean (SEM).
Reduction of NP-40
NP‑40 is a detergent used to suppress protein and compound aggregation and to preserve enzyme activity. While NP‑40 maintained enzyme performance, its concentration critically influenced PAINS‑related artifacts. Specifically, levels below 0.01% or at/above 0.02% were associated with increased apparent PAINS compound activity (data not shown). Optimisation of this buffer component balanced enzyme stability with PAINS‑driven assay interference.
Optimised assay performance
Following buffer optimisation enzyme kinetics were re-evaluated, confirming that assay performance remained robust. The final optimised buffer afforded a much-improved PAINS profile (Figure 5), enabling more reliable data generation by better distinguishing true enzyme inhibitors from compounds that interact with the enzyme in non‑productive or artefactual ways.
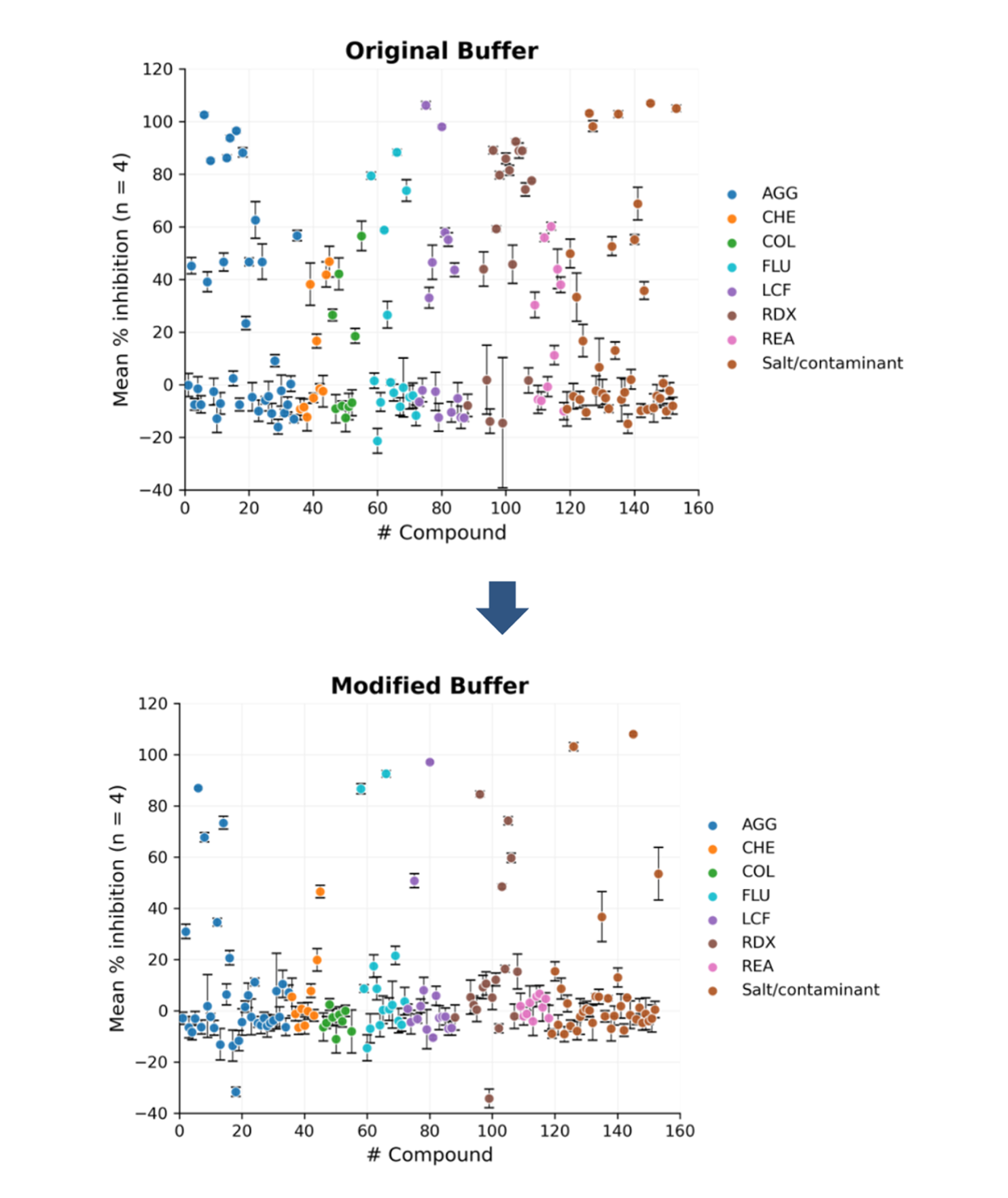
Figure 5. Original assay buffer and modified assay buffer after optimisation using PAINS compound testing. Error bars represent the standard error of the mean (SEM).
By applying this optimisation strategy, we were able to strengthen assay robustness without compromising enzyme activity. The refined buffer conditions were subsequently used to run a successful HTS of 185,000 compounds across two complementary assay formats.
This approach markedly reduced the prevalence of assay-active compounds whose behaviour was driven by non-specific or interference-based mechanisms commonly associated with PAINS chemotypes. The resulting hit lists were then validated using orthogonal biophysical methods (SPR and TSA), which confirmed that the optimised assay conditions effectively filtered out these mechanistically irrelevant actives.
By identifying and mitigating key sources of assay interference upfront, our team enables clients to obtain more reliable screening outcomes, accelerate decision-making, and reduce overall discovery costs.
Learn more about BioAscent’s assay development and screening expertise here.




